Researchers in China observed that cancer patients was 18 out of 1,590 COVID-19 patients (1.132%). The general China population with cancer was 286 out of 100,000 people, 0.286%. It was about 3.95 times of the general China cancer population.
Zong et al. 2021, reported that there are clinical and molecular similarities of COVID-19 and cancers. They summarized the four major signalings pathways at the intersection of COVID-19 and cancers : cytokine, interferon l, AR, and immune checkpoint signaling.
Cytokine Storm (CS) due to the elevated cytokine level is common in COVID-19 and cancers. CS is a key indicator of COVID-19 deterioration. The proinflammatory cytokine IL-6 plays a major role in CS and mediated JAK/STAT signaling. JAK/STAT signaling plays a pivotal role in regulating cell growth, survival, differentiation, motility, and immune responses. JAK/STAT pathway may result in chronic inflammatory conditions or various types of cancers. Therefore drugs targeting the IL-6/JAK/STAT pathway with anticancer effects may be repurposed for the treatment of COVID-19.
IFN-l is common in COVID-19 and cancers. IFNs belongs to a large family of cytokines. IFN-Is bind to the receptor activating JAKs to initiate signal transduction through the JAK/STAT pathway. IFN-I signaling leads to the activation of a multitude of interferon regulatory factors (IRFs) and IFN-stimulated genes (ISGs), thus promoting inflammatory and innate antiviral responses. In cancer biology, IFN-I plays a vital role in inhibiting tumor proliferation and promoting tumor cell senescence and death, and impaired IFN-I signaling is associated with tumor progression. Low levels of IFN-I and ISGs, along with an increase in IL-6 and inflammatory responses, were observed in peripheral blood samples from patients with severe or critical COVID-19. Improper timing of the IFN-I response failed to inhibit viral replication of MERS-CoV and SARS-CoV-2.
ARs (androgen receptors) belong to the superfamily of hormonal nuclear receptors. One of the target genes of ARs is TMPRSS2, which encodes a type II transmembrane protein with serine protease activity. The androgen-regulated TMPRSS2 gene may fuse with the erythroblast transformation-specific (ETS)-related gene ERG, the most common member of the oncogenic ETS family. The mRNA expression level of TMPRSS2 is robustly increased in prostate cancer in response to androgen stimulation. Patients with prostate cancer receiving androgen-deprivation therapy (ADT) were significantly less vulnerable to COVID-19 than those not treated with ADT. TMPRSS2 is the protease that mediates S protein cleavage at the S1/S2 site, is believed to be essential for the interaction of SARS-CoV-2 with ACE2 and cell entry.
Immune checkpoint signaling. ICI (immune checkpoint inhibition) contributes to viral clearance and has been evaluated in the treatment of infectious diseases and it's a promising therapeutic pillar in oncology. Upregulation of immune checkpoint receptors in severe COVID-19 cases is associated with T cell exhaustion and lymphopenia. T cell is one of the immune checkpoint signaling key steps.
TP53 as King of Cancers Gene
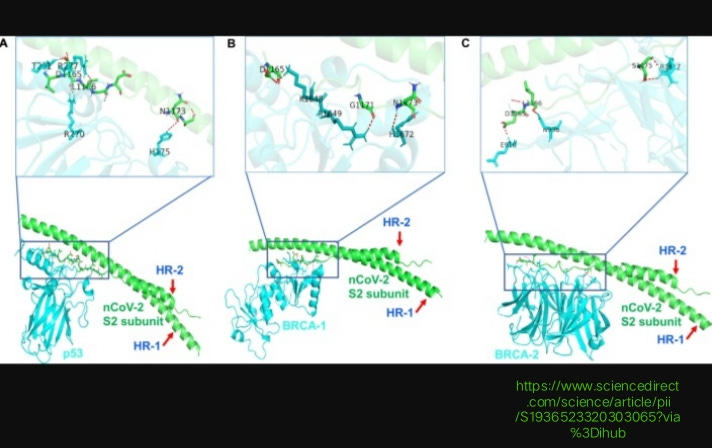
My previous article discussed about the cancer genes hijacked by SARS-CoV-2 proteins. In this article, TP53 in COVID-19 will be explored. Singh et al. 2020, used bioinformatics analytics in investigating interaction of S2 of SARS-COV-2 with TP53 (Tumor Suppressor Proteins P53) and BRCA-1/2 (well known tumor suppressor proteins). BRCA regulates downstream genes in response to numerous cellular stress and are frequently mutated in human cancer. S2 proteins of SARS-CoV-2 mediates the membrane fusion process.
The University of Pittsburgh researchers found that p53, BRCA-1 and BRCA-2 interact with heptic repeat-2 region of S2 subunit through C- terminal domain.
Mutations in the TP53 gene are encountered in about one in every two cases of cancer. Jean C. Perez, 2018, concluded that there is a theoretical method of predicting mutagenic regions of TP53, King of Cancers gene. The mutagenic hotspots and codons referenced from thousands of cases of individual cancers observed (IARC database), correlated with the method. The method is universal and can be applied to the prediction of mutagenic regions of any other gene, protein, human, animal or plant.
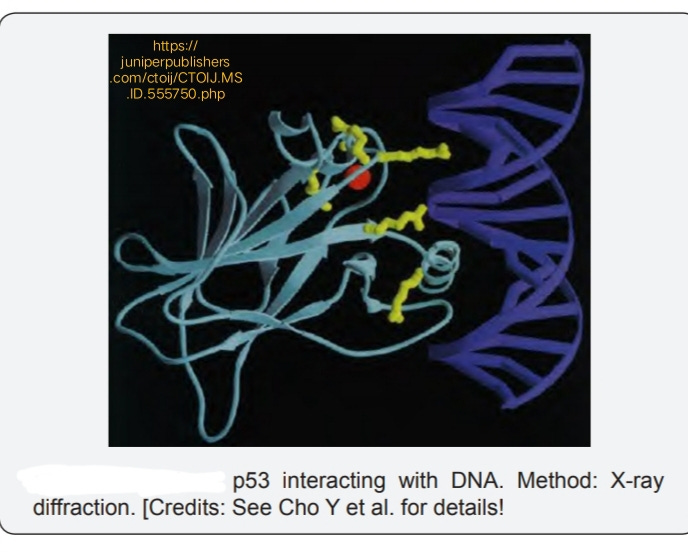
Perez explained that the P53 (TP 53) is a 393 amino acids proteins coming from a long 20kbases length 11 exons gene from the chromosome 17p13. The P53 protein has 5 blocks of highly conserved regions at residues 117-142, 171-181, 234-258 and 270-286. These highly conserved regions coincide with the mutation clusters and HOTSPOTS found in P53 in human cancers, most of which have been found within exons 5 - 8. These mutations have been found to be hightly frequent at the four mutational “hotspots” at codons 175, 245, 248 and 273. Cho Y et al. illustrated the complex interaction provided between P53 and DNA molecule. The DNA (blue) and core domain (turquoise) are shown with the zinc atom (red), with the position of the six hot spot amino acid residues (yellow). Mutations in hot spot amino acids either interfere with protein-DNA contacts, or disrupt integrity of the domain. Thus, all naturally occurring mutations in P53 directly or indirectly affect the interaction of P53 with DNA, demonstrating that sequence-specific DNA binding is central to the normal functioning of P53 as a tumour suppressor.”
Sources:
https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-021-01363-1
https://www.sciencedirect.com/science/article/pii/S1936523320303065?via%3Dihub
https://juniperpublishers.com/ctoij/CTOIJ.MS.ID.555750.php